|
“agents” and collaboration modes) - the best advice I can give is to download it, do the tutorial and test drive it on a bioinformatics task that you regularly conduct - it won't disappoint. The software almost has too many features to comment on in any one review (e.g. Based on the hours of time this program has saved me already I would not even blink at the academic price tag of $133 (USD) - or cheaper for students. The Geneious team has made available a freeware version and a pro version that (for a price) allows you to access more functionality. Moreover, the software is structured in such a way that “new” plugins can be written and incorporated with relative ease. My request to provide a variety of nucleotide colour schemes was implemented within 2 weeks. As is the case with any new software there are a few features that don't work quite as well as you might like - but the good news is that if you write to them they will likely fix it up asap. With the integration of sequence chromatograph viewing and editing of DNA sequences/alignments (new in version 2) Geneious is now firmly embedded in my research practices. If you are looking for a piece of bioinformatics software for teaching purposes look no further than Geneious. Students did a Genbank search, downloaded files, did an alignment and built a phylogenetic tree (with bootstrapping) all in the blink of an eye. 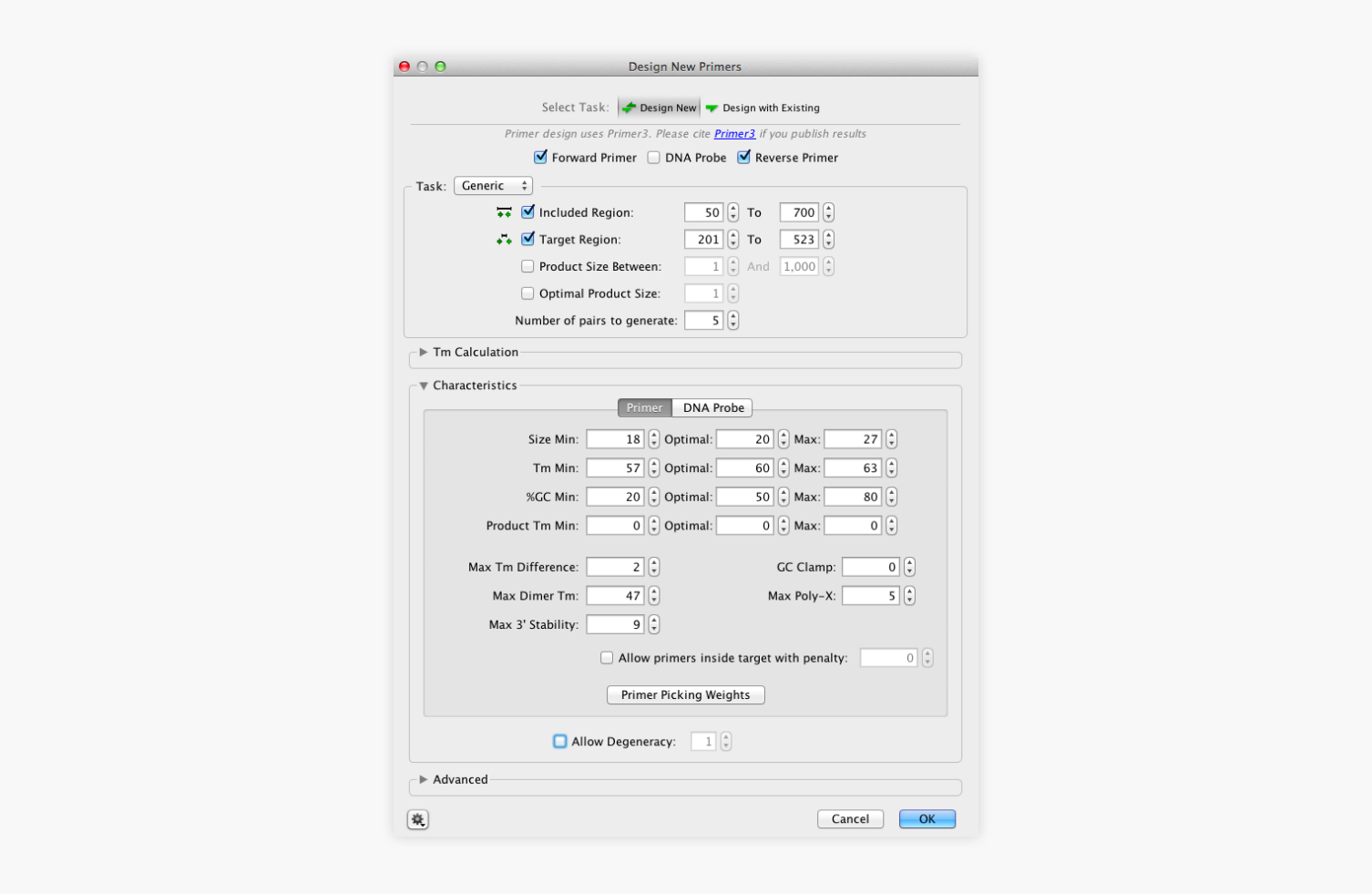
Earlier this year I used the Beta-release in some undergraduate teaching labs - with great success. While the software has some “bell and whistles” for more complex tasks one of its key attributes is a simple user interface. The user interface of Geneious is both attractive and intuitive. The software logo “research in a flash” accurately describe the potential of Geneious - only months after its release it has become the “Swiss army knife” of bioinformatics software. Current and future releases of Geneious (offer a solution to (most) of these frustrations. Chopping and changing between different file formats and at times different operating systems (Mac, PC, Unix/Linux) has been a frustrating and time-consuming exercise. You will then be given the option to delete the pair of that primer at the same time.For a long time scientists who generate or analyse DNA sequences have been using multiple pieces of software to: 1) view chromatographs, 2) trim sequences 3) Blast data 4) align sequences 5) build phylogenetic trees. To delete primers that you don’t want, just select the primer annotation and click the Delete button. If Geneious detects a single Primer Bind annotation with a suitable type IIS site Geneious will use this, and will design a second primer to introduce a compatible opposite orientation restriction site based on rules 4-6. In the case of the reverse primer it will automatically be reverse complemented. This will generate a separate, short sequence document which just contains the primer sequence and the annotation (so it retains all the information on the primer). The best way to save a primer or DNA probe for further testing or use is to select the annotation for that primer and click the Extract button in the sequence viewer. Mismatches between the primer and off-target will be shown in red. An off-target site will be listed if it has no mismatches to the first four 3’ bases of the primer, and less than 10% mismatches with the primer overall. The entire sequence will be searched for off-target sites, even if only a selected region is chosen for primer design. 
These are putative non-specific primer binding sites identified on the sequence that was used for primer design. In Geneious Prime 2019.1 onwards, the primer annotation includes a list of Off-target sites for that primer, including their location and sequence. 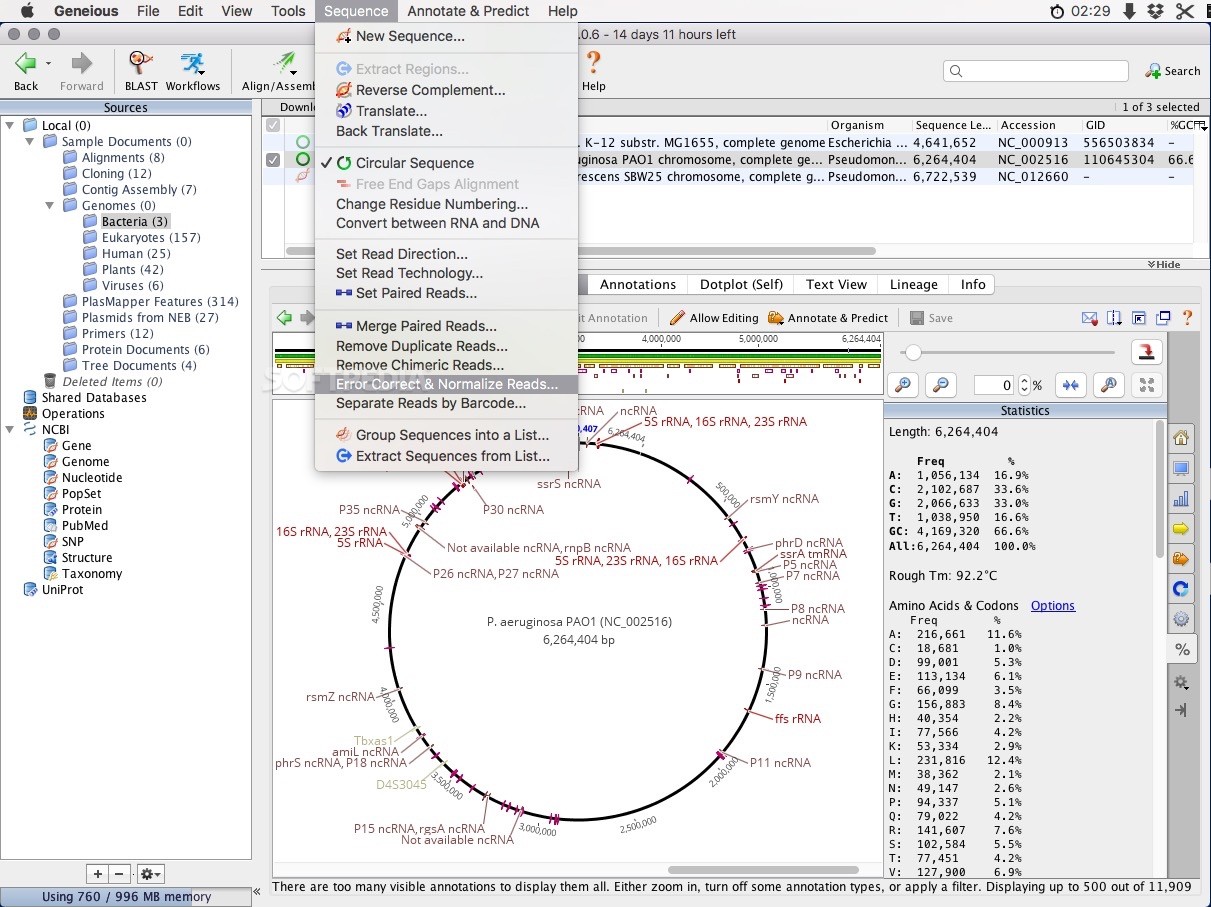
Shows how the values in the Geneious primer annotation map to the original Primer3 values. Alternatively, double clicking on an annotation will display its details in the annotation editing dialog. The information will be presented in a popup box. Primers will be coloured green and probes red.ĭetailed information such as melting point, tendency to form primer-dimers and GC content can be seen by hovering the mouse over the primer annotation. The annotations will be labelled with the base number the primer starts at, followed by either F (forward primer), R (reverse primer), or P (probe). When complete, primers and probes will be added as annotations on the sequences. A progress bar may appear for a short time while the process completes. Once the task and options have been set, click the OK button to design the primers.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
